Importing from an EPrints repository
If your legacy system is an EPrints repository, you will be able to use the haplo_repo_eprints plugin to import your existing data. Install this as a dependency of your root application plugin.
This plugin provides default mappings for data held in an EPrints repository to the Haplo system, and an administrative interface to run, develop, and debug an import.
Import steps
After logging into the application as an administrator:
Add an EPrints admin account to the application’s keychain
To be able to download restricted files the application will need an administrator account to the legacy EPrints instance. Add this to the system keychain as an HTTP/Basic credential with the name “Eprints admin account”, by going to:
Get an export from EPrints in EP3 XML format
If the institution has not been using the internal “comments” fields, this is easiest to do by going to the “advanced search” page, and searching for all items in the year range 1800-2200 (ie. a date range that covers all the contents of the repository). Then choose “EP3 XML” as the export format for the search results.
If internal fields have been used the hosting provider of the EPrints instance will be able to provide an EP3 XML data dump that includes these fields.
Go to /do/hres-repo-eprints/admin
Next, navigate to the (hidden) EPrints administration page, /do/hres-repo-eprints/admin.
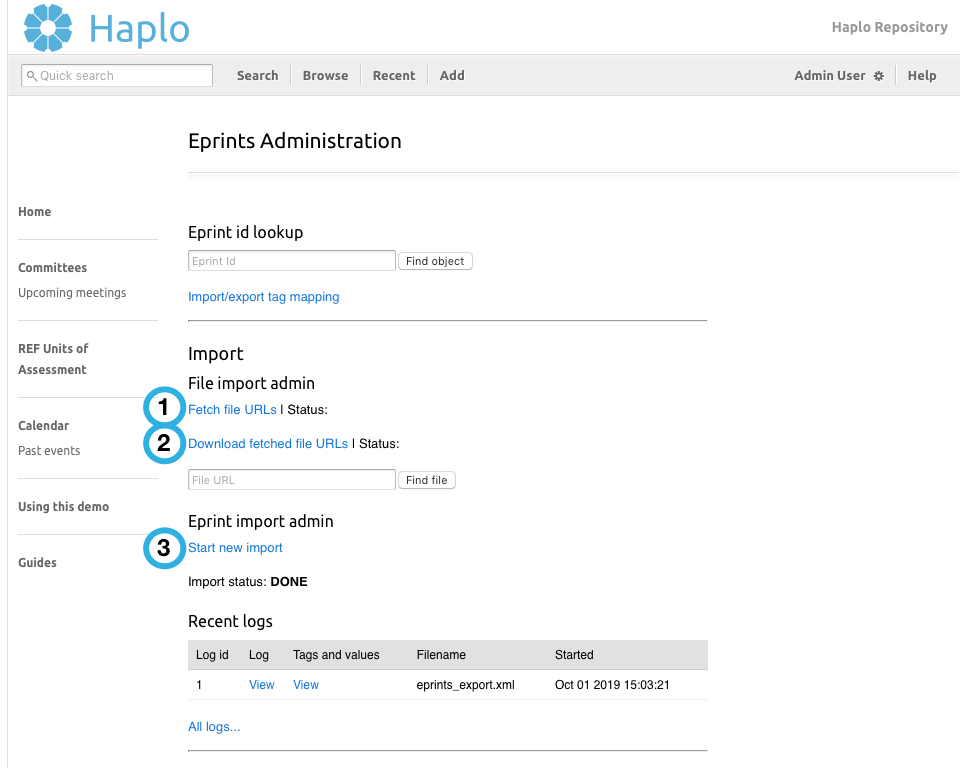
Follow steps 1,2,3 above
Following the links numbered in the admin interface above, upload the exported XML file and confirm each stage of the import process. For convenience while developing you can re-use previously uploaded files without re-uploading them, as they can be quite large.
These steps will, in order:
1) Extract the download URLs for every file in the repository from the uploaded XML
2) Download those files to the Haplo file store, using the “Eprints admin account” keychain credential for restricted items
3) Import all the outputs from the repository, associating the files downloaded in 2) with their metadata records
While each of these stages are running (which may take a couple of hours for larger repositories) the admin interface will display a status of the process.
Logs and troubleshooting
Once the import has completed, a log entry will be added to the Recent logs table. This is a basic JSON document for use by the developer to assess the import and any data cleaning work that may be needed.
The key entries in the JSON log are:
outputsByType |
how many of each type of output have been created |
newObjects |
how many associated objects were created (eg. journal and publisher records) |
eprints |
a lookup of import warnings for specific records, keyed by eprintid |
ERROR:unimported_tags |
a list of any tags encountered in the input XML that have not been either imported or ignored (configured in code) |
The admin interface also includes lookups by eprintid and Haplo Ref to explore how records have been imported into the system, and which tags from EPrints have been ignored.
Since Haplo contains a unique record for each researcher at the institution, there is also a data quality dashboard to report all the items that have not been linked to a known author. This is likely to be a key piece of data cleaning work undertaken during the migration, as items not linked to a researcher’s record will not be listed under their profile in the repository.
Extending the import
As each installation of EPrints is configured differently, it is likely that some customisation of the import will be needed. To do this you should implement a new <clientname>_eprints plugin, and use the service APIs provided by hres_repo_eprints to modify the import process. These will allow you to override, amend, or extend the mappings provided by the generic plugin.
These services, along with more technical details about how the import works in code, are held in the plugin readme.